CLion 2018.2 EAP: argument selection defects inspection
Hi,
Please welcome CLion 2018.2 EAP (build 182.3569.10)!
As usual, a patch-update will be available shortly for those using the previous EAP build, and you can also use Toolbox app or snap packages (in the case of Ubuntu) to get this build. No license is required and the build is free to use, but it will expire within 30 days of the build date.
New inspection: argument selection defects
With this new inspection CLion can detect situations when wrong arguments are passed to a function call (for example, the wrong order of the arguments of the same type). The algorithm relies on various heuristics and some description can be found here. If argument and parameter names are meaningful, the results can be really impressive and useful:

The inspection settings are located in Settings/Preferences | Editor | Inspections | C/C++ | General | Argument selection defects:
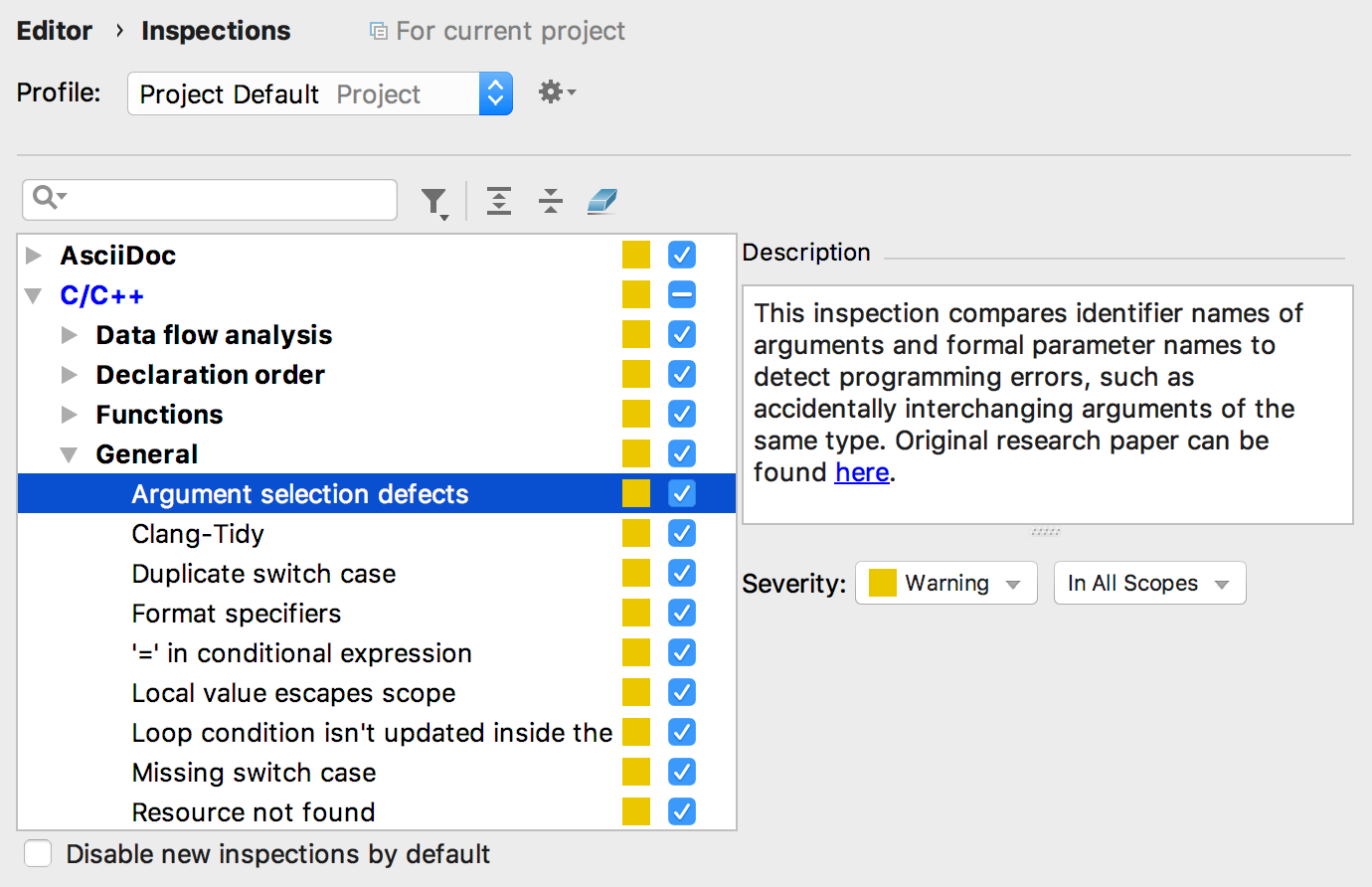
Mind that the inspection is working on top of the experimental clangd-based language engine, so it requires the clang-tidy via a clangd option to be turned on. Among other limitations, note that the inspections is not triggered for the following cases:
- At least one of the parameters have one or two letters in the name (to avoid false positives).
- At least one of the parameters contain “swap/inverse/rotate/backward/flip” substrings.
Other changes
Besides, this build brings the following improvements:
- It fixes the issue with failing build with WSL due to whitespace in path.
- It bundles CMake v3.12.
- If the compilation database for the project you try to open in CLion includes some incorrect compile command, CLion now reports a corresponding error.
It’s also worth mentioning that previous EAP build enabled clangd as an additional C++ language engine in the experimental mode and your feedback and issue reports are very important at this stage.
Full release notes are available by the link.
Your CLion Team
JetBrains
The Drive to Develop